The purpose of this note is to describe the Gaussian hypercontractivity inequality. As an application, we’ll obtain a weaker version of the Hanson–Wright inequality.
The Noise Operator
We begin our discussion with the following question:
Let 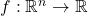 be a function. What happens to
be a function. What happens to 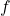 , on average, if we perturb its inputs by a small amount of Gaussian noise?
, on average, if we perturb its inputs by a small amount of Gaussian noise?
Let’s be more specific about our noise model. Let 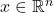 be an input to the function
be an input to the function 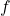 and fix a parameter
and fix a parameter 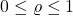 (think of
(think of 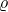 as close to 1). We’ll define the noise corruption of
as close to 1). We’ll define the noise corruption of 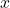 to be
to be
(1) ![Rendered by QuickLaTeX.com \[\tilde{x}_\varrho = \varrho \cdot x + \sqrt{1-\varrho^2} \cdot g, \quad \text{where } g\sim \operatorname{Normal}(0,I). \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-a9a458d4a3787f2f427555dcbe4f377a_l3.png)
Here, 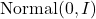 is the standard multivariate Gaussian distribution. In our definition of
is the standard multivariate Gaussian distribution. In our definition of 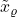 , we both add Gaussian noise
, we both add Gaussian noise 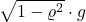 and shrink the vector
and shrink the vector 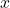 by a factor
by a factor 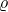 . In particular, we highlight two extreme cases:
. In particular, we highlight two extreme cases:
- No noise. If
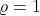 , then there is no noise and
, then there is no noise and 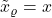 .
.
- All noise. If
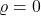 , then there is all noise and
, then there is all noise and 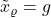 . The influence of the original vector
. The influence of the original vector 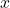 has been washed away completely.
has been washed away completely.
The noise corruption (1) immediately gives rise to the noise operator 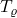 . Let
. Let 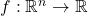 be a function. The noise operator
be a function. The noise operator 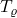 is defined to be:
is defined to be:
(2) ![Rendered by QuickLaTeX.com \[(T_\varrho f)(x) = \expect[f(\tilde{x}_\varrho)] = \expect_{g\sim \operatorname{Normal}(0,I)}[f( \varrho \cdot x + \sqrt{1-\varrho^2}\cdot g)]. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-7d5a50c680b2faee3c89c5a8d21d6496_l3.png)
The noise operator computes the average value of 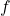 when evaluated at the noisy input
when evaluated at the noisy input 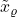 . Observe that the noise operator maps a function
. Observe that the noise operator maps a function 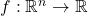 to another function
to another function 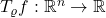 . Going forward, we will write
. Going forward, we will write 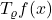 to denote
to denote 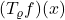 .
.
To understand how the noise operator acts on a function 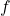 , we can write the expectation in the definition (2) as an integral:
, we can write the expectation in the definition (2) as an integral:
![Rendered by QuickLaTeX.com \[T_\varrho f(x) = \int_{\real^d} f(\varrho x + y) \frac{1}{(2\pi (1-\varrho^2))^{d/2}}\e^{-\frac{|y|^2}{2(1-\varrho^2)}} \, \mathrm{d} y.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c3e346daf451893f22c11a800ebfaf47_l3.png)
Here,
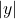
denotes the
(Euclidean) length of
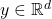
. We see that
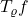
is the
convolution of
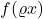
with a Gaussian density. Thus,
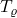
acts to
smooth the function
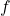
.
See below for an illustration. The red solid curve is a function 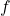 , and the blue dashed curve is
, and the blue dashed curve is 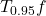 .
.
As we decrease 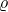 from
from 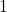 to
to 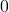 , the function
, the function 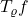 is smoothed more and more. When we finally reach
is smoothed more and more. When we finally reach 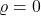 ,
, 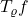 has been smoothed all the way into a constant.
has been smoothed all the way into a constant.
Random Inputs
The noise operator converts a function 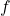 to another function
to another function 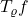 . We can evaluate these two functions at a Gaussian random vector
. We can evaluate these two functions at a Gaussian random vector 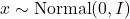 , resulting in two random variables
, resulting in two random variables 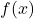 and
and 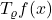 .
.
We can think of 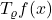 as a modification of the random variable
as a modification of the random variable 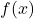 where “a
where “a 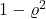 fraction of the variance of
fraction of the variance of 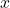 has been averaged out”. We again highlight the two extreme cases:
has been averaged out”. We again highlight the two extreme cases:
- No noise. If
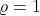 ,
, 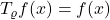 . None of the variance of
. None of the variance of 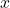 has been averaged out.
has been averaged out.
- All noise. If
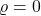 ,
,![Rendered by QuickLaTeX.com T_\varrho f(x) = \expect_{g\sim\operatorname{Normal}(0,I)}[f(g)]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8b96c3890f33c8e43bbeffe16e7ba478_l3.png) is a constant random variable. All of the variance of
is a constant random variable. All of the variance of 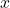 has been averaged out.
has been averaged out.
Just as decreasing 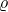 smoothes the function
smoothes the function 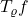 until it reaches a constant function at
until it reaches a constant function at 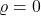 , decreasing
, decreasing 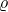 makes the random variable
makes the random variable 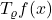 more and more “well-behaved” until it becomes a constant random variable at
more and more “well-behaved” until it becomes a constant random variable at 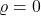 . This “well-behavingness” property of the noise operator is made precise by the Gaussian hypercontractivity theorem.
. This “well-behavingness” property of the noise operator is made precise by the Gaussian hypercontractivity theorem.
Moments and Tails
In order to describe the “well-behavingness” properties of the noise operator, we must answer the question:
How can we measure how well-behaved a random variable is?
There are many answers to this question. For this post, we will quantify the well-behavedness of a random variable by using the 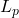 norm.
norm.
The 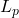 norm of a (
norm of a (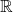 -valued) random variable
-valued) random variable 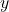 is defined to be
is defined to be
(3) ![Rendered by QuickLaTeX.com \[\norm{y}_p \coloneqq \left( \expect[|y|^p] \right)^{1/p}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-44366ba8d857261afbac3dd53d6772d7_l3.png)
The
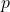
th power of the
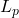
norm
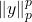
is sometimes known as the
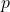 th absolute moment
th absolute moment of
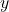
.
The 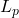 norms of random variables control the tails of a random variable—that is, the probability that a random variable is large in magnitude. A random variables with small tails is typically thought of as a “nice” or “well-behaved” random variable. Random quantities with small tails are usually desirable in applications, as they are more predictable—unlikely to take large values.
norms of random variables control the tails of a random variable—that is, the probability that a random variable is large in magnitude. A random variables with small tails is typically thought of as a “nice” or “well-behaved” random variable. Random quantities with small tails are usually desirable in applications, as they are more predictable—unlikely to take large values.
The connection between tails and 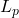 norms can be derived as follows. First, write the tail probability
norms can be derived as follows. First, write the tail probability 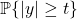 for
for  using
using 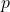 th powers:
th powers:
![Rendered by QuickLaTeX.com \[\prob \{|y| \ge t\} = \prob\{ |y|^p \ge t^p \}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0261832962dbc7be3cd42ee90aad4985_l3.png)
Then, we apply
Markov’s inequality, obtaining
(4) ![Rendered by QuickLaTeX.com \[\prob \{|y| \ge t\} = \prob \{ |y|^p \ge t^p \} \le \frac{\expect [|y|^p]}{t^p} = \frac{\norm{y}_p^p}{t^p}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-82a5fd8d43135158bea4014476475d20_l3.png)
We conclude that a random variable with finite
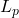
norm (i.e.,
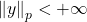
) has tails that decay at at a rate
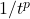
or faster.
Gaussian Contractivity
Before we introduce the Gaussian hypercontractivity theorem, let’s establish a weaker property of the noise operator, contractivity.
Proposition 1 (Gaussian contractivity). Choose a noise level 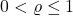 and a power
and a power 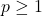 , and let
, and let 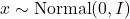 be a Gaussian random vector. Then
be a Gaussian random vector. Then 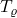 contracts the
contracts the 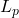 norm of
norm of 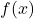 :
:
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_p \le \norm{f(x)}_p.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c0dbc615fb70297848472921d2487389_l3.png)
This result shows that the noise operator makes the random variable 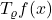 no less nice than
no less nice than 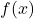 was.
was.
Gaussian contractivity is easy to prove. Begin using the definition of the noise operator (2) and 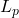 norm (3):
norm (3):
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_p^p = \expect_{x\sim \operatorname{Normal}(0,I)} \left[ \left|\expect_{g\sim \operatorname{Normal}(0,I)}[f(\varrho x + \sqrt{1-\varrho^2}\cdot g)]\right|^p\right]\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-2ad77e2a481ea65ab407d4ea1b97bcf3_l3.png)
Now, we can apply
Jensen’s inequality to the
convex function 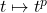
, obtaining
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_p^p \le \expect_{x,g\sim \operatorname{Normal}(0,I)} \left[ \left|f(\varrho x + \sqrt{1-\varrho^2}\cdot g)\right|^p\right].\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-3d906e11f90b7d058484470191771286_l3.png)
Finally, realize that for the independent normal random vectors
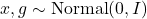
, we have
![Rendered by QuickLaTeX.com \[\varrho x + \sqrt{1-\varrho^2}\cdot g \sim \operatorname{Normal}(0,I).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-099f21f0607675c171a13dfa7f7e9797_l3.png)
Thus,
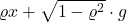
has the same distribution as
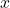
. Thus, using
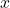
in place of
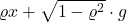
, we obtain
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_p^p \le \expect_{x\sim \operatorname{Normal}(0,I)} \left[ \left|f(x)\right|^p\right] = \norm{f(x)}_p^p.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-7e0cd61fe7d9de9c3599a954b4177ef4_l3.png)
Gaussian contractivity (Proposition 1) is proven.
Gaussian Hypercontractivity
The Gaussian contractivity theorem shows that 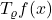 is no less well-behaved than
is no less well-behaved than 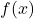 is. In fact,
is. In fact, 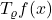 is more well-behaved than
is more well-behaved than 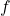 is. This is the content of the Gaussian hypercontractivity theorem:
is. This is the content of the Gaussian hypercontractivity theorem:
Theorem 2 (Gaussian hypercontractivity): Choose a noise level 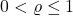 and a power
and a power 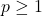 , and let
, and let 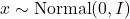 be a Gaussian random vector. Then
be a Gaussian random vector. Then
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_{1+(p-1)/\varrho^2} \le \norm{f(x)}_p.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b7c928cee672afbd1f374bee388fc51f_l3.png)
In particular, for 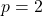 ,
, ![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_{1+\varrho^{-2}} \le \norm{f(x)}_2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-09ccc1018ffef3e9e4259b35417b3c39_l3.png)
We have highlighted the 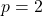 case because it is the most useful in practice.
case because it is the most useful in practice.
This result shows that as we take 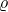 smaller, the random variable
smaller, the random variable 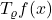 becomes more and more well-behaved, with tails decreasing at a rate
becomes more and more well-behaved, with tails decreasing at a rate
![Rendered by QuickLaTeX.com \[\prob \{ |T_\varrho f(x)| \ge t \} \le \frac{\norm{T_\varrho f(x)}_{1+(p-1)/\varrho^2}^{1+(p-1)/\varrho^2}}{t^{1 + (p-1)/\varrho^2}} \le \frac{\norm{f(x)}_p^{1+(p-1)/\varrho^2}}{t^{1 + (p-1)/\varrho^2}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b88fb5685022862dc4d201ef397412b7_l3.png)
The rate of tail decrease becomes faster and faster as
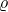
becomes closer to zero.
We will prove the Gaussian hypercontractivity at the bottom of this post. For now, we will focus on applying this result.
Multilinear Polynomials
A multilinear polynomial 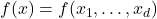 is a multivariate polynomial in the variables
is a multivariate polynomial in the variables 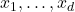 in which none of the variables
in which none of the variables 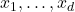 is raised to a power higher than one. So,
is raised to a power higher than one. So,
(5) ![Rendered by QuickLaTeX.com \[1+x_1x_2\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-83d9680ffa5d2d2393e75bfd88937e02_l3.png)
is multilinear, but
![Rendered by QuickLaTeX.com \[1+x_1+x_1x_2^2\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-29f9e247053918faf80d52be03c825b4_l3.png)
is not multilinear (since
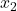
is squared).
For multilinear polynomials, we have the following very powerful corollary of Gaussian hypercontractivity:
Corollary 3 (Absolute moments of a multilinear polynomial of Gaussians). Let 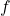 be a multilinear polynomial of degree
be a multilinear polynomial of degree 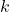 . (That is, at most
. (That is, at most 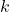 variables
variables 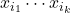 occur in any monomial of
occur in any monomial of 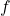 .) Then, for a Gaussian random vector
.) Then, for a Gaussian random vector 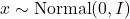 and for all
and for all 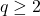 ,
,
![Rendered by QuickLaTeX.com \[\norm{f(x)}_q \le (q-1)^{k/2} \norm{f(x)}_2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c30cae5045c1ed3f76cb40b3777fc260_l3.png)
Let’s prove this corollary. The first observation is that the noise operator has a particularly convenient form when applied to a multilinear polynomial. Let’s test it out on our example (5) from above. For
![Rendered by QuickLaTeX.com \[f(x) = 1+x_1x_2,\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-48daab93e37453735b37017785bf6e4b_l3.png)
we have
![Rendered by QuickLaTeX.com \begin{align*}T_\varrho f(x) &= \expect_{g_1,g_2 \sim \operatorname{Normal}(0,1)} \left[1+ (\varrho x_1 + \sqrt{1-\varrho^2}\cdot g_1)(\varrho x_2 + \sqrt{1-\varrho^2}\cdot g_2)\right].\\&= 1 + \expect[\varrho x_1 + \sqrt{1-\varrho^2}\cdot g_1]\expect[\varrho x_2 + \sqrt{1-\varrho^2}\cdot g_2]\\&= 1+ (\varrho x_1)(\varrho x_2) \\&= f(\varrho x).\end{align*}](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4ef2894b129938470b62a41f8a25cbb8_l3.png)
We see that the expectation applies to each variable separately, resulting in each 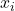 replaced by
replaced by 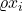 . This trend holds in general:
. This trend holds in general:
Proposition 4 (noise operator on multilinear polynomials). For any multilinear polynomial 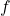 ,
, 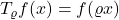 .
.
We can use Proposition 4 to obtain bounds on the 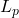 norms of multilinear polynomials of a Gaussian random variable. Indeed, observe that
norms of multilinear polynomials of a Gaussian random variable. Indeed, observe that
![Rendered by QuickLaTeX.com \[f(x) = f(\varrho \cdot x/\varrho) = T_\varrho f(x/\varrho).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-dbff7c144d42223ef42258479652a183_l3.png)
Thus, by Gaussian hypercontractivity, we have
![Rendered by QuickLaTeX.com \[\norm{f(x)}_{1+\varrho^{-2}}=\norm{T_\varrho f(x/\varrho)}_{1+\varrho^{-2}} \le \norm{f(x/\varrho)}_2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-a5da2fc55864dbedbcf6b06f9f0d563c_l3.png)
The final step of our argument will be to compute 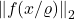 . Write
. Write 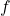 as
as
![Rendered by QuickLaTeX.com \[f(x) = \sum_{i_1,\ldots,i_s} a_{i_1,\ldots,i_s} x_{i_1} \cdots x_{i_s}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-01b58d06d59c8245d6470aae03a2dcb2_l3.png)
Since
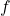
is multilinear,
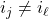
for
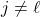
. Since
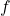
is degree-
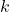
,
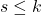
. The multilinear monomials

are
orthonormal with respect to the
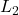
inner product:
![Rendered by QuickLaTeX.com \[\expect[(x_{i_1}\cdots x_{i_s}) \cdot (x_{i_1'}\cdots x_{i_s'})] = \begin{cases} 0 &\text{if } \{i_1,\ldots,i_s\} \ne \{i_1',\ldots,i_{s'}\}, \\1, & \text{otherwise}.\end{cases}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-a48620e82a969b0c929f9b6177a5d0ea_l3.png)
(See if you can see why!) Thus, by the
Pythagorean theorem, we have
![Rendered by QuickLaTeX.com \[\norm{f(x)}_2^2 = \sum_{i_1,\ldots,i_s} a_{i_1,\ldots,i_s}^2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f67f45324bd945aed6f9048c75f07978_l3.png)
Similarly, the coefficients of
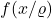
are
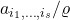
. Thus,
![Rendered by QuickLaTeX.com \[\norm{f(x/\varrho)}_2^2 = \sum_{i_1,\ldots,i_s} \varrho^{-2s} a_{i_1,\ldots,i_s}^2 \le \varrho^{-2k} \sum_{i_1,\ldots,i_s} a_{i_1,\ldots,i_s}^2 = \varrho^{-2k}\norm{f(x)}_2^2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-870a1f48ad3f595f1dcfb8ad67a00e0f_l3.png)
Thus, putting all of the ingredients together, we have
![Rendered by QuickLaTeX.com \[\norm{f(x)}_{1+\varrho^{-2}}=\norm{T_\varrho f(x/\varrho)}_p \le \norm{f(x/\varrho)}_2 \le \varrho^{-k} \norm{f(x)}_2.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4d28f135d2ac1404d6275c2100466b1e_l3.png)
Setting
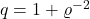
(equivalently
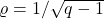
), Corollary 3 follows.
Hanson–Wright Inequality
To see the power of the machinery we have developed, let’s prove a version of the Hanson–Wright inequality.
Theorem 5 (suboptimal Hanson–Wright). Let 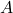 be a symmetric matrix with zero on its diagonal and
be a symmetric matrix with zero on its diagonal and 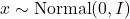 be a Gaussian random vector. Then
be a Gaussian random vector. Then
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \exp\left(- \frac{t}{\sqrt{2}\mathrm{e}\norm{A}_{\rm F}} \right) \quad \text{for } t\ge \sqrt{2}\mathrm{e}\norm{A}_{\rm F}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9236006f1ec332d319b6334c1a30095a_l3.png)
Hanson–Wright has all sorts of applications in computational mathematics and data science. One direct application is to obtain probabilistic error bounds for the error incurred by a stochastic trace estimation formulas.
This version of Hanson–Wright is not perfect. In particular, it does not capture the Bernstein-type tail behavior of the classical Hanson–Wright inequality
![Rendered by QuickLaTeX.com \[\prob\{|x^\top Ax| \ge t\} \le 2\exp \left( -\frac{t^2}{4\norm{A}_{\rm F}^2+4\norm{A}t} \right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d7f7a58dff94bcb23926659eec1dc27c_l3.png)
But our suboptimal Hanson–Wright inequality is still pretty good, and it requires essentially no work to prove using the hypercontractivity machinery. The hypercontractivity technique also generalizes to settings where some of the proofs of Hanson–Wright fail, such as multilinear polynomials of degree higher than two.
Let’s prove our suboptimal Hanson–Wright inequality. Set 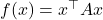 . Since
. Since 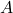 has zero on its diagonal,
has zero on its diagonal, 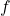 is a multilinear polynomial of degree two in the entries of
is a multilinear polynomial of degree two in the entries of 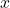 . The random variable
. The random variable 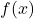 is mean-zero, and a short calculation shows its
is mean-zero, and a short calculation shows its 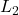 norm is
norm is
![Rendered by QuickLaTeX.com \[\norm{f(x)}_2 = \sqrt{\Var(f(x))} = \sqrt{2} \norm{A}_{\rm F}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-a34d480d366fe0d0606ef8236e673a7f_l3.png)
Thus, by Corollary 3,
(6) ![Rendered by QuickLaTeX.com \[\norm{f(x)}_q \le (q-1) \norm{f(x)}_2 \le \sqrt{2} q \norm{A}_{\rm F} \quad \text{for every } q\ge 2. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d2399e2a6bc3547a4c1950a722ef58da_l3.png)
In fact, since the
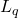
norms are monotone, (6) holds for
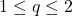
as well. Therefore, the standard tail bound for
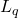
norms (4) gives
(7) ![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \frac{\norm{f(x)}_q^q}{t^q} \le \left( \frac{\sqrt{2}q\norm{A}_{\rm F}}{t} \right)^q\quad \text{for }q\ge 1.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b6c06aa286ad7d5bbac00062d8bf7948_l3.png)
Now, we must optimize the value of 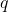 to obtain the sharpest possible bound. To make this optimization more convenient, introduce a parameter
to obtain the sharpest possible bound. To make this optimization more convenient, introduce a parameter
![Rendered by QuickLaTeX.com \[\alpha \coloneqq \frac{\sqrt{2}q\norm{A}_{\rm F}}{t}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-74cc3e742a9613c91e3a6050ea881477_l3.png)
In terms of the
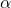
parameter, the bound (7) reads
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \exp\left(- \frac{t}{\sqrt{2}\norm{A}_{\rm F}} \alpha \ln \frac{1}{\alpha} \right) \quad \text{for } t\ge \frac{\sqrt{2}\norm{A}_{\rm F}}{\alpha}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-2d530d1abe13452bcfe5d4a586ee5318_l3.png)
The tail bound is minimized by taking
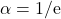
, yielding the claimed result
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \exp\left(- \frac{t}{\sqrt{2}\mathrm{e}\norm{A}_{\rm F}} \right) \quad \text{for } t\ge \sqrt{2}\mathrm{e}\norm{A}_{\rm F}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9236006f1ec332d319b6334c1a30095a_l3.png)
Proof of Gaussian Hypercontractivity
Let’s prove the Gaussian hypercontractivity theorem. For simplicity, we will stick with the 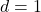 case, but the higher-dimensional generalizations follow along similar lines. The key ingredient will be the Gaussian Jensen inequality, which made a prominent appearance in a previous blog post of mine. Here, we will only need the following version:
case, but the higher-dimensional generalizations follow along similar lines. The key ingredient will be the Gaussian Jensen inequality, which made a prominent appearance in a previous blog post of mine. Here, we will only need the following version:
Theorem 6 (Gaussian Jensen). Let 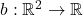 be a twice differentiable function and let
be a twice differentiable function and let 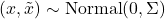 be jointly Gaussian random variables with covariance matrix
be jointly Gaussian random variables with covariance matrix 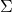 . Then
. Then
(8) ![Rendered by QuickLaTeX.com \[b(\expect[h_1(x)], \expect[h_2(\tilde{x})]) \ge \expect [b(h_1(x),h_2(\tilde{x}))]\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0bffa2a925f66d777f89afb4927d2317_l3.png)
holds for all test functions 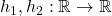 if, and only if,
if, and only if, (9) ![Rendered by QuickLaTeX.com \[\Sigma \circ \nabla^2 b \quad\text{is negative semidefinite on all of $\real^2$}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f2b0499e1b65730027eca4ca776978db_l3.png)
Here,  denotes the entrywise product of matrices and
denotes the entrywise product of matrices and 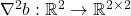 is the Hessian matrix of the function
is the Hessian matrix of the function 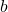 .
.
To me, this proof of Gaussian hypercontractivity using Gaussian Jensen (adapted from Paata Ivanishvili‘s excellent post) is amazing. First, we reformulate the Gaussian hypercontractivity property a couple of times using some functional analysis tricks. Then we do a short calculation, invoke Gaussian Jensen, and the theorem is proved, almost as if by magic.
Part 1: Tricks
Let’s begin with “tricks” part of the argument.
Trick 1. To prove Gaussian hypercontractivity holds for all functions 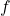 , it is sufficient to prove for all nonnegative functions
, it is sufficient to prove for all nonnegative functions 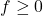 .
.
Indeed, suppose Gaussian hypercontractivity holds for all nonnegative functions 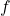 . Then, for any function
. Then, for any function 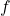 , apply Jensen’s inequality to conclude
, apply Jensen’s inequality to conclude
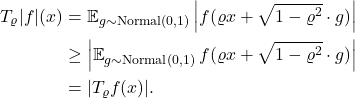
Thus, assuming hypercontractivity holds for the nonnegative function 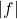 , we have
, we have
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_{1+(p-1)/\varrho^2} \le \norm{T_\varrho |f|(x)}_{1+(p-1)/\varrho^2} \le \norm{|f|(x)}_p = \norm{f}_p.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-bc53021db84adf63753c81b86dc7dd7f_l3.png)
Thus, the conclusion of the hypercontractivity theorem holds for
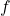
as well, and the Trick 1 is proven.
Trick 2. To prove Gaussian hypercontractivity for all 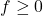 , it is sufficient to prove the following “bilinearized” Gaussian hypercontractivity result:
, it is sufficient to prove the following “bilinearized” Gaussian hypercontractivity result:
![Rendered by QuickLaTeX.com \[\expect[g(x) \cdot T_\varrho f(x)]\le \norm{g(x)}_{q'} \norm{f(x)}_p\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-635e6125f7c6897bee0fdb39ec23147e_l3.png)
holds for all 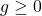 with
with 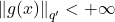 . Here,
. Here, 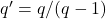 is the Hölder conjugate to
is the Hölder conjugate to 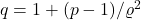 .
.
Indeed, this follows from the dual characterization of the norm of 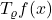 :
:
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_q = \sup_{\substack{\norm{g(x)} < +\infty \\ g\ge 0}} \frac{\expect[g(x) \cdot T_\varrho f(x)]}{\norm{g(x)}_{q'}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d90023ec0671c90d4b5b46f7edf16374_l3.png)
Trick 2 is proven.
Trick 3. Let 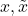 be a pair of standard Gaussian random variables with correlation
be a pair of standard Gaussian random variables with correlation 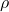 . Then the bilinearized Gaussian hypercontractivity statement is equivalent to
. Then the bilinearized Gaussian hypercontractivity statement is equivalent to
![Rendered by QuickLaTeX.com \[\expect[g(x) f(\tilde{x})]\le (\expect[(g(x)^{q'})])^{1/q'} (\expect[(f(\tilde{x})^{p})])^{1/p}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-fb4614c78266ca91ec0df9d50a073784_l3.png)
Indeed, define 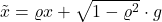 for the random variable in the definition of the noise operator
for the random variable in the definition of the noise operator 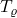 . The random variable
. The random variable 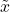 is standard Gaussian and has correlation
is standard Gaussian and has correlation 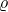 with
with 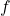 , concluding the proof of Trick 3.
, concluding the proof of Trick 3.
Finally, we apply a change of variables as our last trick:
Trick 4. Make the change of variables 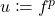 and
and 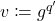 , yielding the final equivalent version of Gaussian hypercontractivity:
, yielding the final equivalent version of Gaussian hypercontractivity:
![Rendered by QuickLaTeX.com \[\expect[v(x)^{1/q'} u(\tilde{x})^{1/p}]\le (\expect[v(x)])^{1/q'} (\expect[u(\tilde{x}))])^{1/p}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-22bcbc15be00ee01a0d33b8f61597707_l3.png)
for all functions 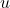 and
and  (in the appropriate spaces).
(in the appropriate spaces).
Part 2: Calculation
We recognize this fourth equivalent version of Gaussian hypercontractivity as the conclusion (8) to Gaussian Jensen with
![Rendered by QuickLaTeX.com \[b(u,v) = u^{1/p}v^{1/q'}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-22fc749a1125afbfa747d3fd2f8265d0_l3.png)
. Thus, to prove Gaussian hypercontractivity, we just need to check the hypothesis (9) of the Gaussian Jensen inequality (Theorem 6).
We now enter the calculation part of the proof. First, we compute the Hessian of 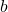 :
:
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v) = u^{1/p}v^{1/q'}\cdot\begin{bmatrix} - \frac{1}{pp'} u^{-2} & \frac{1}{pq'} u^{-1}v^{-1} \\ \frac{1}{pq'} u^{-1}v^{-1} & - \frac{1}{qq'} v^{-2}\end{bmatrix}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c8be66a18ac99521a2bf99265ec9f5e1_l3.png)
We have written
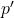
for the
Hölder conjugate to
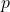
. By Gaussian Jensen, to prove Gaussian hypercontractivity, it suffices to show that
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v)\circ \twobytwo{1}{\varrho}{\varrho}{1}= u^{1/p}v^{1/q'}\cdot\begin{bmatrix} - \frac{1}{pp'} u^{-2} & \frac{\varrho}{pq'} u^{-1}v^{-1} \\ \frac{\varrho}{pq'} u^{-1}v^{-1} & - \frac{1}{qq'} v^{-2}\end{bmatrix}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-18833758e69322596753a770bad28258_l3.png)
is negative semidefinite for all
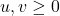
. There are a few ways we can make our lives easier. Write this matrix as
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v)\circ \twobytwo{1}{\varrho}{\varrho}{1}= u^{1/p}v^{1/q'}\cdot B^\top\begin{bmatrix} - \frac{p}{p'} & \varrho \\ \varrho & - \frac{q'}{q} \end{bmatrix}B \quad \text{for } B = \operatorname{diag}(p^{-1}u^{-1},(q')^{-1}v^{-1}).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4a8bc4c34d4bc535576fa960b4ef612d_l3.png)
Scaling
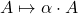
by nonnegative
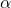
and conjugation
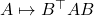
both preserve negative semidefiniteness, so it is sufficient to prove
![Rendered by QuickLaTeX.com \[H = \begin{bmatrix} - \frac{p}{p'} & \varrho \\ \varrho & - \frac{q'}{q} \end{bmatrix} \quad \text{is negative semidefinite}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f68417fc42aa107dba992f15c5e65b4a_l3.png)
Since the diagonal entries of
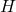
are negative, at least one of
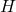
‘s eigenvalues is negative. Therefore, to prove
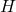
is negative semidefinite, we can prove that its determinant (= product of its eigenvalues) is nonnegative. We compute
![Rendered by QuickLaTeX.com \[\det H = \frac{pq'}{p'q} - \varrho^2 .\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-571f5f1397c668c88b8426c34e2d52ad_l3.png)
Now, just plug in the values for
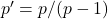
,
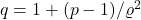
,
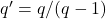
:
![Rendered by QuickLaTeX.com \[\det H = \frac{pq'}{p'q} - \varrho^2 = \frac{p-1}{q-1} - \varrho^2 = \frac{p-1}{(p-1)/\varrho^2} - \varrho^2 = 0.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-dc40f6da9dd8637b19397b29d27e85ed_l3.png)
Thus,
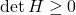
. We conclude
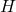
is negative semidefinite, proving the Gaussian hypercontractivity theorem.
![]() be vectors, assemble the matrix
be vectors, assemble the matrix ![]() , and form the Gram matrix
, and form the Gram matrix![]()
![]()
![]()
be strictly positive numbers. Then the inverse of their average is no bigger than the average of their inverses:
![]() into a
into a ![]() positive semidefinite matrix
positive semidefinite matrix ![]() . Taking the average of all such matrices, we observe that
. Taking the average of all such matrices, we observe that ![]()
![Rendered by QuickLaTeX.com \[\det(A) = \left(\frac{1}{n} \sum_{i=1}^n x_i\right) \left(\frac{1}{n} \sum_{i=1}^n \frac{1}{x_i}\right) - 1 \ge 0.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-e1e6cd521eaf4f49c17bab509cf71cd2_l3.png)
![Rendered by QuickLaTeX.com \[\left( \frac{1}{n} \sum_{i=1}^n x_i \right)^{-1} \le \frac{1}{n} \sum_{i=1}^n \frac{1}{x_i}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-db85840bf998e96281f1c2605ca740c5_l3.png)
![Rendered by QuickLaTeX.com \[\hat{A} &\gets \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right)^\top \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d691d56d7580112fdd2d2db8eb4cd4aa_l3.png)
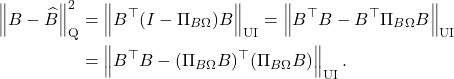
![Rendered by QuickLaTeX.com \[\kappa_{\rm dem} = \frac{\norm{A}_{\rm F}}{\sigma_{\rm min}(A)} = \sqrt{\sum_i \left(\frac{\sigma_i(A)}{\sigma_{\rm min}(A)}\right)^2} \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0a01bce740f28deced2d1a8b4e6e1002_l3.png)
![Rendered by QuickLaTeX.com \[D \coloneqq \diag\left( \sqrt{\frac{p_j}{p_j^{\rm RK}}} : j =1,\ldots,n\right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-bc930770f7b4e20afffe903a4ab372ba_l3.png)
![Rendered by QuickLaTeX.com \[x_{t+1} = \left( I - \frac{a_{i_t}^{\vphantom{\top}}a_{i_t}^\top}{\norm{a_{i_t}}^2} \right)x_t + \frac{b_{i_t}a_{i_t}}{\norm{a_{i_t}}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-cccfb54762bb990499c96331e7019309_l3.png)
![Rendered by QuickLaTeX.com \begin{align*}\expect_{i_t}[x_{t+1}] &= \sum_{j=1}^n \left[\left( I - \frac{a_j^{\vphantom{\top}} a_j^\top}{\norm{a_j}^2} \right)x_t + \frac{b_ja_j}{\norm{a_j}^2}\right] \prob\{i_t=j\}\\ &=\sum_{j=1}^n \left[\left( \frac{\norm{a_j}^2}{\norm{A}_{\rm F}^2}I - \frac{a_j^{\vphantom{\top}} a_j^\top}{\norm{A}_{\rm F}^2} \right)x_t + \frac{b_ja_j}{\norm{A}_{\rm F}^2}\right].\end{align*}](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0a7f7b3bfad51b7094348db94d2d4254_l3.png)
![Rendered by QuickLaTeX.com \[\expect_{i_t}[x_{t+1}] = \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right) x_t + \frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-de4ded17daf001dde92c26025e9c14f3_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_{t+1}] = \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right) \expect[x_t] + \frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-24c5bfe6c1ea70293055deb68a3df117_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] = \left[\sum_{i=0}^{t-1} \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-eb47c96091b1187b53c318589b696693_l3.png)
![Rendered by QuickLaTeX.com \[\left(\frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^{-1} = \sum_{i=0}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8d1be156303c168d2de83dbde76e44d6_l3.png)
![Rendered by QuickLaTeX.com \[\left(\frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^{-1} = \sum_{i=0}^{t-1} \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i + \sum_{i=t}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-08f4b1a705439315fe0fc038e059d01c_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] = x_\star - \left[\sum_{i=t}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d973041dba49ba471bbfb6b3d4c2b463_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] - x_\star = - \left(I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^t \left[\sum_{i=0}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8238e65e7b7df2013504f2a6f71799e6_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] - x_\star = - \left(I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^t x_\star.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-3cbe72663f8ef13c762213b5592bb5d2_l3.png)
![Rendered by QuickLaTeX.com \[\norm{\expect[x_t] - x_\star}^2 \le \norm{I - \frac{A^\top A}{\norm{A}_{\rm F}^2}}^{2t} \norm{x}_\star^2. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-e663370119bd02900da0aa4f7b6faf03_l3.png)
![Rendered by QuickLaTeX.com \[I - \frac{A^\top A}{\norm{A}_{\rm F}^2} = I - V \cdot\frac{\Sigma^2}{\norm{A}_{\rm F}^2} \cdot V^\top = V \left( I - \frac{\Sigma^2}{\norm{A}_{\rm F}^2}\right)V^\top.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-468566feb9b39f9e0dd616b47f324ad5_l3.png)
![Rendered by QuickLaTeX.com \[\xi_{\overline{q}}(\theta) \le \xi_{g^\top \overline{A} g}(\theta) \le \frac{\theta^2 \norm{\overline{A}}_{\rm F}^2}{1 - 2\theta \norm{\overline{A}}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-623e28881a1cb46a4526687b07e28e0e_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{ x^\top A x - \tr(A) \ge t \} \le \exp \left( - \frac{t^2/2}{2 \norm{\overline{A}}_{\rm F}^2 + 2 \overline{\norm{\overline{A}}t}} \right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-16071627e38437191476b04f643b1057_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{ |x^\top A x - \tr(A)| \ge t \} \le 2\exp \left( - \frac{t^2/2}{2 \norm{\overline{A}}_{\rm F}^2 + 2 \overline{\norm{\overline{A}}t}} \right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f19454359ca5d0e68b0207b865db95db_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{ g^\top A g - \tr A \ge t \} \le \exp \left( -\frac{t^2/2}{C\norm{A}_{\rm F}^2 + c t \norm{A}}\right)\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-cb3646b736a9e33403f0175d6bc8e296_l3.png)
![Rendered by QuickLaTeX.com \[\xi_{\overline{\tr}}(\theta) = \sum_{i=1}^m \xi_{x_i^\top Ax_i}(\theta/m) \le m \cdot \frac{(\theta/m)^2 \norm{\overline{A}}_{\rm F}^2}{1 - 2(\theta/m)\norm{\overline{A}}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d6d7f289a105bd06e272223f56ee7080_l3.png)
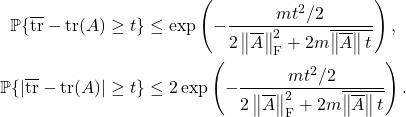
![Rendered by QuickLaTeX.com \[\prob \{\text{any accepted this loop}\} = \sum_{i=1}^n \prob \{\text{$i$ accepted this loop}\} = \frac{\sum_{i=1}^n w_i}{\sum_{i=1}^n \rho_i}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-87004ae4b99269f6917de03d3a4ec86e_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{ |T_\varrho f(x)| \ge t \} \le \frac{\norm{T_\varrho f(x)}_{1+(p-1)/\varrho^2}^{1+(p-1)/\varrho^2}}{t^{1 + (p-1)/\varrho^2}} \le \frac{\norm{f(x)}_p^{1+(p-1)/\varrho^2}}{t^{1 + (p-1)/\varrho^2}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b88fb5685022862dc4d201ef397412b7_l3.png)
![Rendered by QuickLaTeX.com \begin{align*}T_\varrho f(x) &= \expect_{g_1,g_2 \sim \operatorname{Normal}(0,1)} \left[1+ (\varrho x_1 + \sqrt{1-\varrho^2}\cdot g_1)(\varrho x_2 + \sqrt{1-\varrho^2}\cdot g_2)\right].\\&= 1 + \expect[\varrho x_1 + \sqrt{1-\varrho^2}\cdot g_1]\expect[\varrho x_2 + \sqrt{1-\varrho^2}\cdot g_2]\\&= 1+ (\varrho x_1)(\varrho x_2) \\&= f(\varrho x).\end{align*}](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4ef2894b129938470b62a41f8a25cbb8_l3.png)
![Rendered by QuickLaTeX.com \[\expect[(x_{i_1}\cdots x_{i_s}) \cdot (x_{i_1'}\cdots x_{i_s'})] = \begin{cases} 0 &\text{if } \{i_1,\ldots,i_s\} \ne \{i_1',\ldots,i_{s'}\}, \\1, & \text{otherwise}.\end{cases}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-a48620e82a969b0c929f9b6177a5d0ea_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \exp\left(- \frac{t}{\sqrt{2}\mathrm{e}\norm{A}_{\rm F}} \right) \quad \text{for } t\ge \sqrt{2}\mathrm{e}\norm{A}_{\rm F}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9236006f1ec332d319b6334c1a30095a_l3.png)
![Rendered by QuickLaTeX.com \[\prob\{|x^\top Ax| \ge t\} \le 2\exp \left( -\frac{t^2}{4\norm{A}_{\rm F}^2+4\norm{A}t} \right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d7f7a58dff94bcb23926659eec1dc27c_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \frac{\norm{f(x)}_q^q}{t^q} \le \left( \frac{\sqrt{2}q\norm{A}_{\rm F}}{t} \right)^q\quad \text{for }q\ge 1.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b6c06aa286ad7d5bbac00062d8bf7948_l3.png)
![Rendered by QuickLaTeX.com \[\prob \{|x^\top A x| \ge t \} \le \exp\left(- \frac{t}{\sqrt{2}\norm{A}_{\rm F}} \alpha \ln \frac{1}{\alpha} \right) \quad \text{for } t\ge \frac{\sqrt{2}\norm{A}_{\rm F}}{\alpha}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-2d530d1abe13452bcfe5d4a586ee5318_l3.png)
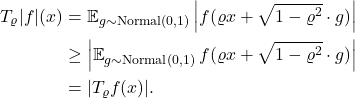
![Rendered by QuickLaTeX.com \[\norm{T_\varrho f(x)}_q = \sup_{\substack{\norm{g(x)} < +\infty \\ g\ge 0}} \frac{\expect[g(x) \cdot T_\varrho f(x)]}{\norm{g(x)}_{q'}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d90023ec0671c90d4b5b46f7edf16374_l3.png)
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v) = u^{1/p}v^{1/q'}\cdot\begin{bmatrix} - \frac{1}{pp'} u^{-2} & \frac{1}{pq'} u^{-1}v^{-1} \\ \frac{1}{pq'} u^{-1}v^{-1} & - \frac{1}{qq'} v^{-2}\end{bmatrix}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c8be66a18ac99521a2bf99265ec9f5e1_l3.png)
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v)\circ \twobytwo{1}{\varrho}{\varrho}{1}= u^{1/p}v^{1/q'}\cdot\begin{bmatrix} - \frac{1}{pp'} u^{-2} & \frac{\varrho}{pq'} u^{-1}v^{-1} \\ \frac{\varrho}{pq'} u^{-1}v^{-1} & - \frac{1}{qq'} v^{-2}\end{bmatrix}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-18833758e69322596753a770bad28258_l3.png)
![Rendered by QuickLaTeX.com \[\nabla^2 b(u,v)\circ \twobytwo{1}{\varrho}{\varrho}{1}= u^{1/p}v^{1/q'}\cdot B^\top\begin{bmatrix} - \frac{p}{p'} & \varrho \\ \varrho & - \frac{q'}{q} \end{bmatrix}B \quad \text{for } B = \operatorname{diag}(p^{-1}u^{-1},(q')^{-1}v^{-1}).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4a8bc4c34d4bc535576fa960b4ef612d_l3.png)
![Rendered by QuickLaTeX.com \[H = \begin{bmatrix} - \frac{p}{p'} & \varrho \\ \varrho & - \frac{q'}{q} \end{bmatrix} \quad \text{is negative semidefinite}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f68417fc42aa107dba992f15c5e65b4a_l3.png)