I am delighted to share that my paper Randomly pivoted Cholesky: Randomly pivoted Cholesky: Practical approximation of a kernel matrix with few entry evaluations, joint with Yifan Chen, Joel Tropp, and Robert Webber, has been published online in Communications on Pure and Applied Mathematics. To celebrate this occasion, I want to share one of my favorite tricks in the design of low-rank approximation algorithms, which I will call the Gram correspondence.
Projection Approximation and Nyström Approximation
When we construct a low-rank approximation to a matrix, the type of approximation we use is typically dictated by the size and properties of the matrix. For a rectangular matrix 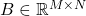 , one of the standard techniques is the standard randomized SVD algorithm:
, one of the standard techniques is the standard randomized SVD algorithm:
- Generate a Gaussian random matrix
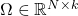 .
.
- Form the product
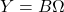 .
.
- Compute an (economy-size) QR decomposition
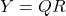 .
.
- Evaluate the SVD
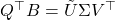 .
.
- Output the low-rank approximation
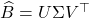 for
for 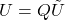 .
.
Succinctly, the output of the randomized SVD is given by the formula 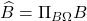 , where
, where 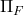 denotes the orthogonal projector onto the column span of a matrix
denotes the orthogonal projector onto the column span of a matrix 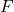 . This motivates the general definition:
. This motivates the general definition:
Definition (projection approximation): Given a test matrix 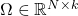 , the projection approximation to the matrix
, the projection approximation to the matrix  is
is 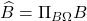 .
.
The class of projection approximations is much richer than merely the approximation constructed by the standard randomized SVD. Indeed, low-rank approximations computed by randomized subspace iteration, randomized block Krylov iteration, column-pivoted QR decompositions, etc. all fit under the umbrella of projection approximations.
Many matrices in applications have additional structure such as symmetry or sparsity, and it can be valuable to make use of low-rank approximations that take advantage of those properties. An especially important type of structure is positive semidefiniteness. For our purposes, a positive semidefinite matrix 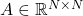 is one that is symmetric and possesses nonnegative eigenvalues, and we will use abbreviate “positive semidefinite” as “psd”. Psd matrices arise in applications as covariance matrices, kernel matrices, and as discretizations of certain differential and integral operators. Further, any rectangular matrix
is one that is symmetric and possesses nonnegative eigenvalues, and we will use abbreviate “positive semidefinite” as “psd”. Psd matrices arise in applications as covariance matrices, kernel matrices, and as discretizations of certain differential and integral operators. Further, any rectangular matrix 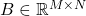 gives rise to its psd Gram matrix
gives rise to its psd Gram matrix 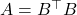 ; we will have much more to say about Gram matrices below.
; we will have much more to say about Gram matrices below.
To compute a low-rank approximation to a psd matrix, the preferred format as usually the Nyström approximation:
Definition (Nyström approximation): Given a test matrix 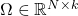 , the Nyström approximation is
, the Nyström approximation is
![Rendered by QuickLaTeX.com \[\hat{A} \coloneqq (A\Omega) (\Omega^\top A \Omega)^{-1}(A\Omega)^\top.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-345eb2a18a6428fbbe9cfa2d305a3026_l3.png)
As discussed in a previous post, the general class of Nyström approximations includes many useful specializations depending on how the matrix 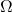 is selected.
is selected.
The Gram Correspondence
The Gram correspondence is a connection between projection approximation and Nyström approximation:
The Gram correspondence: Let 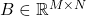 be any rectangular matrix and consider the Gram matrix
be any rectangular matrix and consider the Gram matrix 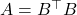 . Fix a test matrix
. Fix a test matrix 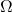 , and define the Nyström approximation
, and define the Nyström approximation 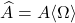 to
to  and the projection approximation
and the projection approximation 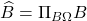 to
to  . Then
. Then
![Rendered by QuickLaTeX.com \[\hat{A} = \smash{\hat{B}}^\top \hat{B}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-70de4cbe941b96a1dc7655f240027afd_l3.png)
That is, the Gram matrix 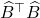 of the projection approximation
of the projection approximation 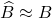 is the Nyström approximation
is the Nyström approximation 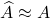 .
.
As we will see, this correspondence has many implications:
- Special cases. The correspondence contains several important facts as special cases. Examples include the equivalence between the randomized SVD and single-pass Nyström approximation and the equivalence of (partial) column-pivoted QR factorization and (partial) pivoted Cholesky decomposition.
- Algorithm design. Since Nyström approximation and projection approximations are closely related, one can often interconvert algorithms for one type of approximation into the other. These conversions can lead one to discover new algorithms. We will provide a historical answer with the discovery of randomly pivoted Cholesky.
- Error bounds. The Gram correspondence is immensely helpful in the analysis of algorithms, as it makes error bounds for projection approximations and Nyström approximations easily derivable from each other.
For those interested, we give a short proof of the Gram correspondence below.
Proof of Gram correspondence
The Gram correspondence is straightforward to derive, so let’s do so before moving on. Because
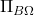
is a projection matrix, it satisfies
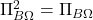
. Thus,
![Rendered by QuickLaTeX.com \[\hat{B}^\top \hat{B} = B^\top \Pi_{B\Omega}^2B = B^\top \Pi_{B\Omega}B.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-fd1a575e449aa0954d4f62e109d824fe_l3.png)
The projector
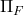
has the formula
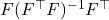
. Using this formula, we obtain
![Rendered by QuickLaTeX.com \[\hat{B}^\top \hat{B} = B^\top B\Omega (\Omega^\top B^\top B \Omega)^{-1} \Omega^\top B^\top B.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f3a9aa2fc08a78569f1745222d2565d5_l3.png)
This is precisely the formula for the Nyström approximation
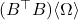
, confirming the Gram correspondence.
Aside: Gram Square Roots
Before we go forward, let me highlight a point of potential conclusion and introduce some helpful terminology. When thinking about algorithms for a psd matrix  , it will often to be helpful to conjure a matrix
, it will often to be helpful to conjure a matrix  for which
for which 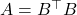 . Given a psd matrix
. Given a psd matrix  , there is always a matrix
, there is always a matrix  for which
for which 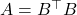 , but this
, but this  is not unique. Indeed, if
is not unique. Indeed, if 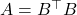 , then we have the infinite family of decompositions
, then we have the infinite family of decompositions 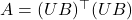 generated by every matrix
generated by every matrix 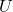 with orthonormal columns. This motivates the following definition:
with orthonormal columns. This motivates the following definition:
Definition (Gram square root): A matrix  is a Gram square root for
is a Gram square root for  if
if 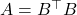 .
.
A Gram square root  for
for  need not be a square root in the traditional sense that
need not be a square root in the traditional sense that 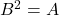 . Indeed, a Gram square root
. Indeed, a Gram square root  can be rectangular, so
can be rectangular, so 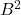 need not even be defined. Using the Gram square root terminology, the Gram correspondence can be written more succinctly:
need not even be defined. Using the Gram square root terminology, the Gram correspondence can be written more succinctly:
Gram correspondence (concise): If  is any Gram square root of
is any Gram square root of  , then the projection approximation
, then the projection approximation 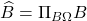 is a Gram square root of the Nyström approximation
is a Gram square root of the Nyström approximation 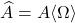 .
.
Examples of the Gram Correspondence
Before going further, let us see a couple of explicit examples of the Gram correspondence.
Randomized SVD and Single-Pass Nyström Approximation
The canonical example of the Gram correspondence is the equivalence of the randomized SVD algorithm and the single-pass Nyström approximation. Let  be a Gram square root for a psd matrix
be a Gram square root for a psd matrix  . The randomized SVD of
. The randomized SVD of  is given by the following steps, which we saw in the introduction:
is given by the following steps, which we saw in the introduction:
- Generate a Gaussian random matrix
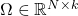 .
.
- Form the product
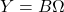 .
.
- Compute a QR decomposition
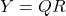 .
.
- Evaluate the SVD
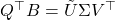 .
.
- Output the low-rank approximation
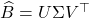 for
for 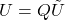 .
.
Now, imagine taking the same Gaussian matrix 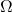 from the randomized SVD, and use it to compute the Nyström approximation
from the randomized SVD, and use it to compute the Nyström approximation 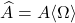 . Notice that this Nyström approximation can be computed in a single pass over the matrix
. Notice that this Nyström approximation can be computed in a single pass over the matrix  . Namely, use a single pass over
. Namely, use a single pass over  to compute
to compute 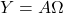 and use following formula:
and use following formula:
![Rendered by QuickLaTeX.com \[\hat{A} = Y (\Omega^\top Y)^{-1} Y^\top.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-439e3ea265c77f2fcf84c6b310f185aa_l3.png)
For this reason, we call
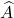
the single-pass Nyström approximation.
The Gram correspondence tells us that the randomized SVD and single-pass Nyström approximation are closely related, in the sense that 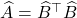 . The randomized SVD approximation
. The randomized SVD approximation 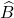 is a Gram square root of the single-pass Nyström approximation
is a Gram square root of the single-pass Nyström approximation 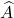 .
.
Column-Pivoted QR Factorization and Pivoted Cholesky Decomposition
A more surprising consequence of the Gram correspondence is the connection between low-rank approximations produced by partial column-pivoted QR factorization of a rectangular matrix  and a partial pivoted Cholesky decomposition of
and a partial pivoted Cholesky decomposition of  . Let’s begin by describing these two approximation methods.
. Let’s begin by describing these two approximation methods.
Let’s begin with pivoted partial Cholesky decomposition. Let 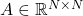 be a psd matrix, and initialize the zero approximation
be a psd matrix, and initialize the zero approximation 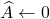 . For
. For 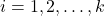 , perform the following steps:
, perform the following steps:
- Choose a column index
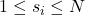 . These indices are referred to as pivot indices or, more simply, pivots.
. These indices are referred to as pivot indices or, more simply, pivots.
- Update the low-rank approximation
![Rendered by QuickLaTeX.com \[\hat{A} \gets \hat{A} + \frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-863d08314865254415f4bb93e143faa9_l3.png)
- Update the matrix
![Rendered by QuickLaTeX.com \[A \gets A - \frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9c0c04b9e4c6217f257a30771739903c_l3.png)
Here, we are using MATLAB notation to index the matrix  . The output of this procedure is
. The output of this procedure is
![Rendered by QuickLaTeX.com \[\hat{A} = A(:,S) A(S,S)^{-1} A(S,:),\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-bbbddde5089193acc3707f2cc631eddd_l3.png)
where
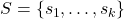
. This type of low-rank approximation is known as a
column Nyström approximation and is a special case of the general Nyström approximation with test matrix
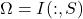
equal to a subset of columns of the identity matrix
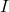
. For an explanation of why this procedure is called a “pivoted partial Cholesky decomposition” and the relation to the usual notion of Cholesky decomposition, see
this previous post of mine.
Given the Gram correspondence, we expect that this pivoted partial Cholesky procedure for computing a Nyström approximation to a psd matrix  should have an analog for computing a projection approximation to a rectangular matrix
should have an analog for computing a projection approximation to a rectangular matrix  . This analog is given by the partial column-pivoted QR factorization, which produces a low-rank approximation according as follows. Let
. This analog is given by the partial column-pivoted QR factorization, which produces a low-rank approximation according as follows. Let 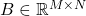 be a psd matrix, and initialize the zero approximation
be a psd matrix, and initialize the zero approximation 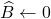 . For
. For 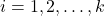 , perform the following steps:
, perform the following steps:
- Choose a pivot index
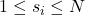 .
.
- Update the low-rank approximation by adding the projection of
 onto the selected column:
onto the selected column: 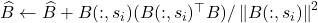 .
.
- Update the matrix by removing the projection of
 onto the selected column:
onto the selected column: 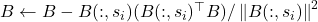 .
.
The output of this procedure is the column projection approximation 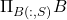 , which is an example of the general projection approximation with
, which is an example of the general projection approximation with 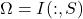 .
.
The pivoted partial Cholesky and QR factorizations are very traditional ways of computing a low-rank approximation in numerical linear algebra. The Gram correspondence tells us immediately that these two approaches are closely related:
Let  be a Gram square root of
be a Gram square root of  . Compute a column projection approximation
. Compute a column projection approximation 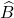 to
to  using a partially column-pivoted QR factorization with pivots
using a partially column-pivoted QR factorization with pivots 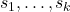 , and compute a column Nyström approximation
, and compute a column Nyström approximation 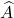 to
to  using a partial pivoted Cholesky decomposition with the same set of pivots
using a partial pivoted Cholesky decomposition with the same set of pivots 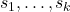 . Then
. Then 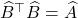 .
.
I find this remarkable: the equivalence of the randomized SVD and single-pass randomized Nyström approximation and the equivalence of (partial pivoted) Cholesky and QR factorizations are both consequences of the same general principle, the Gram correspondence.
Using the Gram Correspondence to Discover Algorithms
The Gram correspondence is more than just an interesting way of connecting existing types of low-rank approximations: It can be used to discover new algorithms. We can illustrate this with an example, the randomly pivoted Cholesky algorithm.
Background: The Randomly Pivoted QR Algorithm
In 2006, Deshpande, Rademacher, Vempala, and Wang discovered an algorithm that they called adaptive sampling for computing a projection approximation to a rectangular matrix 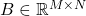 . Beginning from the trivial initial approximation
. Beginning from the trivial initial approximation 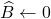 , algorithm proceeds as follows for
, algorithm proceeds as follows for 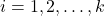 :
:
- Choose a column index
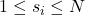 randomly according to the squared column norm distribution:
randomly according to the squared column norm distribution: ![Rendered by QuickLaTeX.com \[\prob\{s_i = j\} = \frac{\norm{B(:,j)}^2}{\norm{B}_{\rm F}^2} \quad \text{for } j=1,2,\ldots,k.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-69b8b3115bb28983cf8ffde1e706bbad_l3.png)
- Update the low-rank approximation by adding the projection of
 onto the selected column:
onto the selected column: 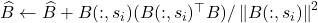 .
.
- Update the matrix by removing the projection of
 onto the selected column:
onto the selected column: 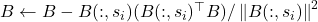 .
.
Given our discussion above, we recognize this as an example of partial column-pivoted QR factorization with a particular randomized procedure for selecting the pivots 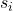 . Therefore, it can also be convenient to call this algorithm randomly pivoted QR.
. Therefore, it can also be convenient to call this algorithm randomly pivoted QR.
Randomly pivoted QR is a nice algorithm for rectangular low-rank approximation. Each step requires a full pass over the matrix and 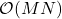 operations, so the full procedure requires
operations, so the full procedure requires 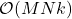 operations. This makes the cost of the algorithm similar to other methods for rectangular low-rank approximation such as the randomized SVD, but it has the advantage that it computes a column projection approximation. However, randomly pivoted QR is not a particularly effective algorithm for computing a low-rank approximation to a psd matrix, since—as we shall see—there are faster procedures available.
operations. This makes the cost of the algorithm similar to other methods for rectangular low-rank approximation such as the randomized SVD, but it has the advantage that it computes a column projection approximation. However, randomly pivoted QR is not a particularly effective algorithm for computing a low-rank approximation to a psd matrix, since—as we shall see—there are faster procedures available.
Following the Gram correspondence, we expect there should be an analog of the randomly pivoted QR algorithm for computing a low-rank approximation of a psd matrix. That algorithm, which we call randomly pivoted Cholesky, is derived from randomly pivoted QR in the following optional section:
Derivation of Randomly Pivoted Cholesky Algorithm
Let
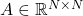
be a psd matrix. For conceptual purposes, let us also consider a Gram square root
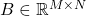
of

; this matrix

will only be a conceptual device that we will use to help us derive the appropriate algorithm, not something that will be required to run the algorithm itself. Let us now walk through the steps of the randomly pivoted QR algorithm on

, and see how they lead to a
randomly pivoted Cholesky algorithm for computing a low-rank approximation to

.
Step 1 of randomly pivoted QR. Randomly pivoted QR begins by drawing a random pivot 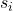 according to the rule
according to the rule
![Rendered by QuickLaTeX.com \[\prob\{s_i = j\} = \frac{\norm{B(:,j)}^2}{\norm{B}_{\rm F}^2} \quad \text{for } j=1,2,\ldots,k.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-69b8b3115bb28983cf8ffde1e706bbad_l3.png)
Since
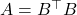
, we may compute
![Rendered by QuickLaTeX.com \[\norm{B(:,j)}^2 = B(:,j)^\top B(:,j) = (B^\top B)(j,j) = A(j,j).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8cc2adf4cfa7b83d8d6cf2f459be5e18_l3.png)
The squared column norms of

are the diagonal entries of the matrix

. Similarly,
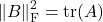
. Therefore, we can write the probability distribution for the random pivot
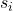
using only the matrix

as
![Rendered by QuickLaTeX.com \[\prob\{s_i = j\} = \frac{A(j,j)}{\tr A} \quad \text{for } j=1,2,\ldots,N.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-203e590f41ae6a725f52f179d8c00c7c_l3.png)
Step 2 of randomly pivoted QR. The randomly pivoted QR update rule is 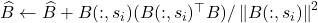 . Therefore, the update rule for
. Therefore, the update rule for 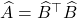 is
is
![Rendered by QuickLaTeX.com \[\hat{A} &\gets \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right)^\top \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c91dc552516b91b718159b2a4a798ed3_l3.png)
After a short derivation,, this simplifies to the update rule
![Rendered by QuickLaTeX.com \[\hat{A} \gets \hat{A} +\frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f5b4e7c853586b6bd3ae380ec4c9f9fb_l3.png)
Two remarkable things have happened. First, we have obtained an update rule for
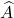
that depends only on the psd matrices
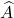
and

, and all occurences of the Gram square roots
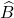
and

have vanished. Second, this update rule is exactly the update rule for a partial Cholesky decomposition.
Step 3 of randomly pivoted QR. Using a similar derivation to step 2, we update an update rule for  :
:
![Rendered by QuickLaTeX.com \[A \gets A -\frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c5812e0143a84e8e070ef048c409df97_l3.png)
This completes the derivation.
Randomly Pivoted Cholesky
That derivation was a bit complicated, so let’s summarize. We can with the randomly pivoted QR algorithm for computing a projection approximation to a rectangular matrix  , and we used it to derive a randomly pivoted Cholesky algorithm for computing a column Nyström approximation to a psd matrix
, and we used it to derive a randomly pivoted Cholesky algorithm for computing a column Nyström approximation to a psd matrix  . Removing the cruft of the derivation, this algorithm is very simple to state:
. Removing the cruft of the derivation, this algorithm is very simple to state:
- Choose a column index
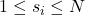 randomly according to the diagonal distribution:
randomly according to the diagonal distribution: ![Rendered by QuickLaTeX.com \[\prob\{s_i = j\} = \frac{A(j,j)}{\tr(A)} \quad \text{for } j=1,2,\ldots,k.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-284bb185bf576727c1adf2287ce110e7_l3.png)
- Update the low-rank approximation
![Rendered by QuickLaTeX.com \[\hat{A} \gets \hat{A} + \frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-863d08314865254415f4bb93e143faa9_l3.png)
- Update the matrix
![Rendered by QuickLaTeX.com \[A \gets A - \frac{A(:,s_i)A(s_i,:)}{A(s_i,s_i)}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9c0c04b9e4c6217f257a30771739903c_l3.png)
I find this to be remarkably cool. We started with a neat algorithm (randomly pivoted QR) for approximating rectangular matrices, and we used the Gram correspondence to derive a different randomly pivoted Cholesky algorithm for psd matrix approximation!
And randomly pivoted Cholesky allows us to pull a cool trick that we couldn’t with randomly pivoted QR. Observe that the randomly pivoted Cholesky algorithm only ever interacts with the residual matrix  through the selected pivot columns
through the selected pivot columns 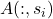 and through the diagonal entries
and through the diagonal entries 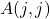 . Therefore, we can derive an optimized version of the randomly pivoted Cholesky algorithm that only reads
. Therefore, we can derive an optimized version of the randomly pivoted Cholesky algorithm that only reads 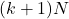 entries of the matrix
entries of the matrix  (
(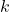 columns plus the diagonal) and requires only
columns plus the diagonal) and requires only 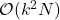 operations! See our paper for details.
operations! See our paper for details.
So we started with the randomly pivoted QR algorithm, which requires 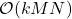 operations, and we used it to derive the randomly pivoted Cholesky algorithm that runs in
operations, and we used it to derive the randomly pivoted Cholesky algorithm that runs in 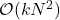 operations. Let’s make this concrete with some specific numbers. Setting
operations. Let’s make this concrete with some specific numbers. Setting 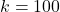 and
and 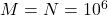 , randomly pivoted QR requires roughly
, randomly pivoted QR requires roughly 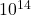 (100 trillion) operations and randomly pivoted Cholesky requires roughly
(100 trillion) operations and randomly pivoted Cholesky requires roughly 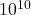 (10 billion) operations, a factor of 10,000 smaller operation count!
(10 billion) operations, a factor of 10,000 smaller operation count!
Randomly pivoted Cholesky has an interesting history. It appears to have first appeared in print in the work of Musco and Woodruff (2017), who used the Gram correspondence to derive the algorithm from randomly pivoted QR. It is remarkable that it took a full 11 years after Deshpande and co-author’s original work on randomly pivoted QR for randomly pivoted Cholesky to be discovered. Even after Musco and Woodruff’s paper, the algorithm appears to have largely been overlooked for computation in practice, and I am unaware of any paper documenting computational experiments with randomly pivoted Cholesky before our paper was initially released in 2022. Our paper reexamined the randomly pivoted Cholesky procedure, providing computational experiments comparing it to alternatives and using it for scientific machine learning applications, and provided new error bounds.
Other Examples of Algorithm Design by the Gram Correspondence
The Gram correspondence gives a powerful tool for inventing new algorithms or showing equivalence between existing algorithms. There are many additional examples, such as block (and rejection sampling-accelerated) versions of randomly pivoted QR/Cholesky, Nyström versions of randomized block Krylov iteration, and column projection/Nyström approximations generated by determinantal point process sampling. Indeed, if you invent a new low-rank approximation algorithm, it is always worth checking: Using the Gram correspondence, can you get another algorithm for free?
Transference of Error Bounds
The Gram correspondence also allows us to transfer error bounds between projection approximations and Nyström approximations. Let’s begin with the simplest manifestation of this principle:
Transference of Error Bounds I: Let  be a Gram square root of
be a Gram square root of  , and let
, and let 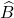 and
and 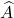 be the corresponding projection and Nyström approximations generated using the same test matrix
be the corresponding projection and Nyström approximations generated using the same test matrix 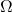 . Then
. Then
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}_{\rm F}^2 = \tr(A - \hat{A}).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-1e2e33e007787db8ac81b5521705652c_l3.png)
This shows that the Frobenius norm error of a projection approximation and the trace error of a Nyström approximation are the same. Using this result, we can immediately transfer error bounds for projection approximations to Nyström approximations and visa versa. For instance, in this previous post, we proved the following error bound for the rank-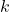 randomized SVD (with a Gaussian test matrix
randomized SVD (with a Gaussian test matrix 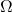 ):
):
![Rendered by QuickLaTeX.com \[\expect \norm{B - \hat{B}}_{\rm F}^2 \le \min_{r\le k-2} \left(1 + \frac{r}{k-(r+1)} \right) \norm{B - \lowrank{B}_r}_{\rm F}^2,\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-1323433654642dc38b1335a77469553e_l3.png)
Here,
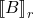
denotes the
best rank-

approximation to

. This result shows that the error of the rank-
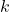
randomized SVD is comparable to the best rank-

approximation for any
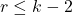
. Using the transference principle, we immediately obtain a corresponding bound for rank-
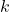
Nyström approximation (with a Gaussian test matrix
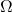
):
![Rendered by QuickLaTeX.com \[\expect \tr(A - \hat{A}) \le \min_{r\le k-2} \left(1 + \frac{r}{k-(r+1)} \right) \tr(A - \lowrank{A}_r).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-5407074bdbcdd0a7918d506ac605b306_l3.png)
With no additional work, we have converted an error bound for the randomized SVD to an error bound for single-pass randomized Nyström approximation!
For those interested, one can extend the transference of error bounds to general unitarily invariant norms. See the following optional section for details:
Transference of Error Bounds for General Unitarily Invariant Norms
We begin by recalling the notions of (quadratic) unitarily invariant norms; see
this section of a previous post for a longer refresher. A norm
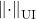
is
unitarily invariant if
![Rendered by QuickLaTeX.com \[\norm{UCV}_{\rm UI} = \norm{C}_{\rm UI}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-6911f19fc91e0e3556bdb09361531e6c_l3.png)
for every matrix
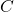
and orthogonal matrices
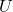
and
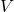
.
Examples of unitarily invariant norms include the nuclear norm
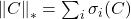
, the Frobenius norm
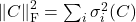
, and the spectral norm
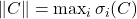
. (It is not coincidental that all these unitarily invariant norms can be expressed only in terms of the
singular values 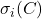
of the matrix
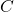
.) Associated to every unitarily invariant norm it its associated
quadratic unitarily invariant norm, defined as
![Rendered by QuickLaTeX.com \[\norm{C}_{\rm Q}^2 = \norm{C^\top C}_{\rm UI} = \norm{CC^\top}_{\rm UI}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f627dafc92609fec4bfd79b7449235f3_l3.png)
For example,
- The quadratic unitarily invariant norm associated to the nuclear norm
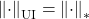 is the Frobenius norm
is the Frobenius norm 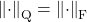 .
.
- The spectral norm is its own associated quadratic unitarily invariant norm
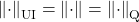 .
.
This leads us to the more general version of the transference principle:
Transference of Error Bounds II: Let  be a Gram square root of
be a Gram square root of  , and let
, and let 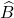 and
and 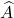 be the corresponding projection and Nyström approximations generated using the same test matrix
be the corresponding projection and Nyström approximations generated using the same test matrix 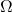 . Then
. Then
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}_{\rm Q}^2 = \norm{A - \hat{A}}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-2fc3e0434bc399a8e268f4009b4ebe52_l3.png)
for every unitarily invariant norm 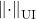 and its associated quadratic unitarily invariant norm
and its associated quadratic unitarily invariant norm 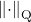 .
.
This version of the principle is more general, since the nuclear norm of a psd matrix is the trace, whence
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}_{\rm F}^2 = \norm{A - \hat{A}}_* = \tr(A - \hat{A}).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-72b523183dfe27ed53b2d5e9fe5ae56a_l3.png)
We now also have more examples. For instance, for the spectral norm, we have
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}^2 = \norm{A - \hat{A}}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b8376bea9952904e84ad58071c48665d_l3.png)
Let us quickly prove the transference of error principle. Write the error 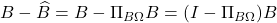 . Thus,
. Thus,
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}_{\rm Q}^2 = \norm{(I-\Pi_{B\Omega})B}_{\rm Q}^2 = \norm{B^\top (I-\Pi_{B\Omega})^2B}_{\rm UI}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-7a4c680324355da1e53b0d72d315f48c_l3.png)
The orthogonal projections
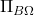
and
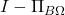
are equal to their own square, so
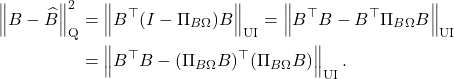
Finally, write
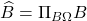
and use the Gram correspondence
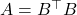
,
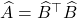
to conclude
![Rendered by QuickLaTeX.com \[\norm{B - \hat{B}}_{\rm Q}^2 = \norm{A - \hat{A}}_{\rm UI},\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-9e9f3b1f06460a7883813043f8cd7e71_l3.png)
as desired.
Conclusion
This has been a long post about a simple idea. Many of the low-rank approximation algorithms in the literature output a special type of approximation, either a projection approximation or a Nyström approximation. As this post has shown, these two types of approximation are equivalent, with algorithms and error bounds for one type of approximation transferring immediately to the other format. This equivalence is a powerful tool for the algorithm designer, allowing us to discover new algorithms from old ones, like randomly pivoted Cholesky from randomly pivoted QR.
![Rendered by QuickLaTeX.com \[\kappa_{\rm dem}(A) = \frac{\norm{A}_{\rm F}}{\sigma_{\rm min}(A)} = \sqrt{\sum_i \left(\frac{\sigma_i(A)}{\sigma_{\rm min}(A)}\right)^2} \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-237843884b53186512cc573dd3d38f7e_l3.png)
![Rendered by QuickLaTeX.com \[\operatorname{sign}(x) = \begin{cases} 1, & x > 0, \\ 0, & x = 0, \\ -1, & x < 0. \end{cases}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-22ae590796d5ac3d80f8159c041a1b90_l3.png)
![Rendered by QuickLaTeX.com \[\langle \varphi_i ,\varphi_j\rangle = \begin{cases}1, & i = j, \\0, & i \ne j.\end{cases}\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-d58150ec2dbdf33688af181314034dc8_l3.png)
![Rendered by QuickLaTeX.com \[\mathcal{E}(f) = \frac{1}{2} \sum_{i,j=1}^m (f(i)-f(j))^2 \pi_iP_{ij}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-e44591ad4cd3c8ec6e25af5f1b4d240c_l3.png)
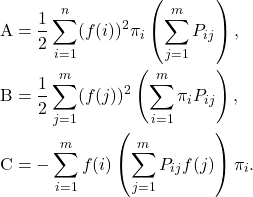
![Rendered by QuickLaTeX.com \[{\rm B} = \frac{1}{2} \sum_{j=1}^m (f(j))^2 \left(\sum_{i=1}^m \pi_j P_{ji} \right) = \frac{1}{2} \sum_{j=1}^m (f(j))^2 \pi_j = \frac{1}{2} \langle f, f\rangle.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-f3b32d0fdd24cdd0d0fd87ed788f8a43_l3.png)
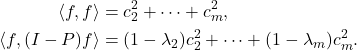
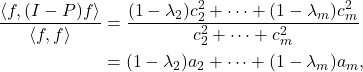
![Rendered by QuickLaTeX.com \[x^\top (A\circ M)x = \sum_{i,j=1}^n x_i (A\circ M)_{ij} x_j = \sum_{i,j=1}^n x_i A_{ij} M_{ij} x_j.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-ca498529303f53df914b6597120a87ba_l3.png)
![Rendered by QuickLaTeX.com \[x^\top (A\circ M)x = \sum_{i,j=1}^n x_i A_{ij} x_j M_{ji} = \tr(\operatorname{diag}(x) A \operatorname{diag}(x) M).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0a90d263429e7e03a177a6cf648480ca_l3.png)
![Rendered by QuickLaTeX.com \[\left( \frac{1}{n} \sum_{i=1}^n x_i \right)^{-1} \le \frac{1}{n} \sum_{i=1}^n \frac{1}{x_i}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-ff75b76509202932a39a33e6748ba18c_l3.png)
![Rendered by QuickLaTeX.com \[\det(A) = \left(\frac{1}{n} \sum_{i=1}^n x_i\right) \left(\frac{1}{n} \sum_{i=1}^n \frac{1}{x_i}\right) - 1 \ge 0.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8c87d66274b7a87984791af241e45f9e_l3.png)
![Rendered by QuickLaTeX.com \[\hat{A} &\gets \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right)^\top \left(\hat{B} + \frac{B(:,s_i) (B(:,s_i)^\top B)}{\norm{B(:,s_i)}^2}\right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c91dc552516b91b718159b2a4a798ed3_l3.png)
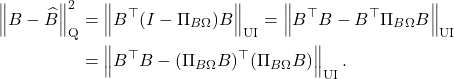
![Rendered by QuickLaTeX.com \[\kappa_{\rm dem} = \frac{\norm{A}_{\rm F}}{\sigma_{\rm min}(A)} = \sqrt{\sum_i \left(\frac{\sigma_i(A)}{\sigma_{\rm min}(A)}\right)^2} \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-6fbef84dde9dc3a326a898ad4e6cc0a2_l3.png)
![Rendered by QuickLaTeX.com \[D \coloneqq \diag\left( \sqrt{\frac{p_j}{p_j^{\rm RK}}} : j =1,\ldots,n\right).\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-c4a48ae205003d974f3d9c22f05b4971_l3.png)
![Rendered by QuickLaTeX.com \[x_{t+1} = \left( I - \frac{a_{i_t}^{\vphantom{\top}}a_{i_t}^\top}{\norm{a_{i_t}}^2} \right)x_t + \frac{b_{i_t}a_{i_t}}{\norm{a_{i_t}}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-ebdcce812d0667315ce96a9a10076256_l3.png)
![Rendered by QuickLaTeX.com \begin{align*}\expect_{i_t}[x_{t+1}] &= \sum_{j=1}^n \left[\left( I - \frac{a_j^{\vphantom{\top}} a_j^\top}{\norm{a_j}^2} \right)x_t + \frac{b_ja_j}{\norm{a_j}^2}\right] \prob\{i_t=j\}\\ &=\sum_{j=1}^n \left[\left( \frac{\norm{a_j}^2}{\norm{A}_{\rm F}^2}I - \frac{a_j^{\vphantom{\top}} a_j^\top}{\norm{A}_{\rm F}^2} \right)x_t + \frac{b_ja_j}{\norm{A}_{\rm F}^2}\right].\end{align*}](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-6313b1e410f036a4151bb5c5f6cb89c8_l3.png)
![Rendered by QuickLaTeX.com \[\expect_{i_t}[x_{t+1}] = \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right) x_t + \frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-5d568d3ca43156f34ddcb4556ac18c68_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_{t+1}] = \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right) \expect[x_t] + \frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-0ac2ab733dcde2fc5c33d3d1b668523c_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] = \left[\sum_{i=0}^{t-1} \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-320f299e0726caf09895b9ce10722c65_l3.png)
![Rendered by QuickLaTeX.com \[\left(\frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^{-1} = \sum_{i=0}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-6b584a1bef1b7a5776457988bb29d3b7_l3.png)
![Rendered by QuickLaTeX.com \[\left(\frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^{-1} = \sum_{i=0}^{t-1} \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i + \sum_{i=t}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-000ddcb9b9d9b6ee0fc32c784f7348c7_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] = x_\star - \left[\sum_{i=t}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-6400bfb8abb56d715838ee2386b5ebd2_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] - x_\star = - \left(I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^t \left[\sum_{i=0}^\infty \left( I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^i \right]\cdot\frac{A^\top b}{\norm{A}_{\rm F}^2}.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-8d61b65cfb176614048ae1b5976d47bf_l3.png)
![Rendered by QuickLaTeX.com \[\expect[x_t] - x_\star = - \left(I - \frac{A^\top A}{\norm{A}_{\rm F}^2}\right)^t x_\star.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-4deff646d7b889bd0c37907ea6a64dbb_l3.png)
![Rendered by QuickLaTeX.com \[\norm{\expect[x_t] - x_\star}^2 \le \norm{I - \frac{A^\top A}{\norm{A}_{\rm F}^2}}^{2t} \norm{x}_\star^2. \]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-cd024bd8769d75cbb989c08c3d4834da_l3.png)
![Rendered by QuickLaTeX.com \[I - \frac{A^\top A}{\norm{A}_{\rm F}^2} = I - V \cdot\frac{\Sigma^2}{\norm{A}_{\rm F}^2} \cdot V^\top = V \left( I - \frac{\Sigma^2}{\norm{A}_{\rm F}^2}\right)V^\top.\]](https://www.ethanepperly.com/wp-content/ql-cache/quicklatex.com-b9d13c9f13a4e81778d6d48f25d891e5_l3.png)